VirulentHunter: Deep Learning-Based Virulence Factor Predictor Illuminates Pathogenicity in Diverse Microbial Niches
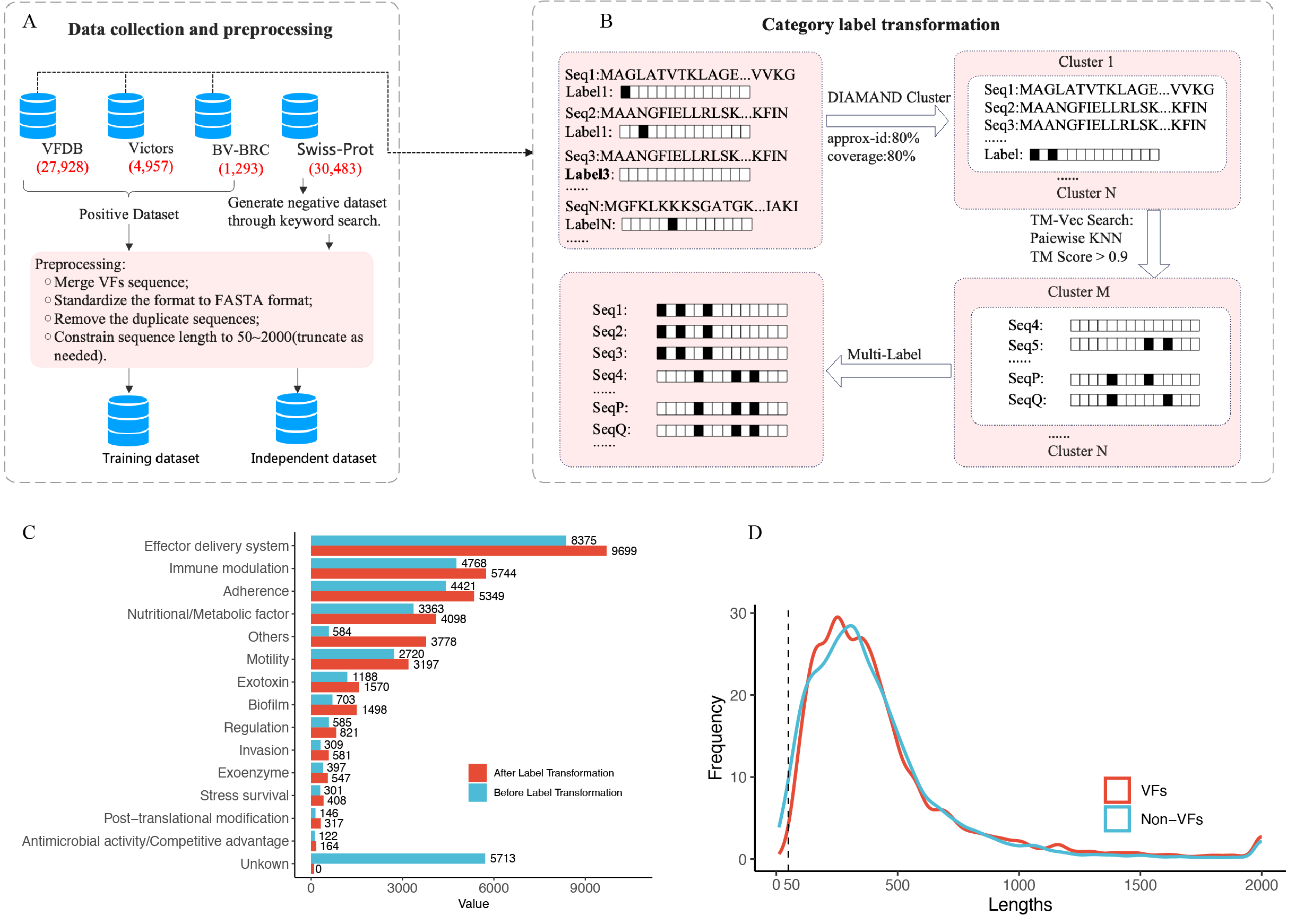
In this study, we focused exclusively on VFs found in bacteria. To do this, we gathered all bacterial VF protein sequences from three public databases, which were VFDB 2022[1], Victors[1] and BV-BRC[3]. Subsequently, these sequences were clustered with CD-HIT[4] v4.8.1, and the duplicates were removed with 100% sequence identity and 80% coverage, yielding 30,483 non-redundant VFs. Because many of the collected VFs lacked category information, we implemented a rigorous label propagation strategy to annotate them using the 14 primary categories defined by the VFDB. Initially, we performed sequence-based clustering using DIAMOND with a sequence identity threshold of 80% and a coverage threshold of 80%. Within each cluster, we assigned all member VFs the combined labels from the union of their existing annotations. To further enhance category assignment, we employed TM-Vec[5], a deep learning tool for structural similarity detection. VFs were clustered using TM-Vec with a threshold of 0.9, and, following the same label propagation strategy, each VF within these clusters was assigned the combined labels from the union of their existing annotations.
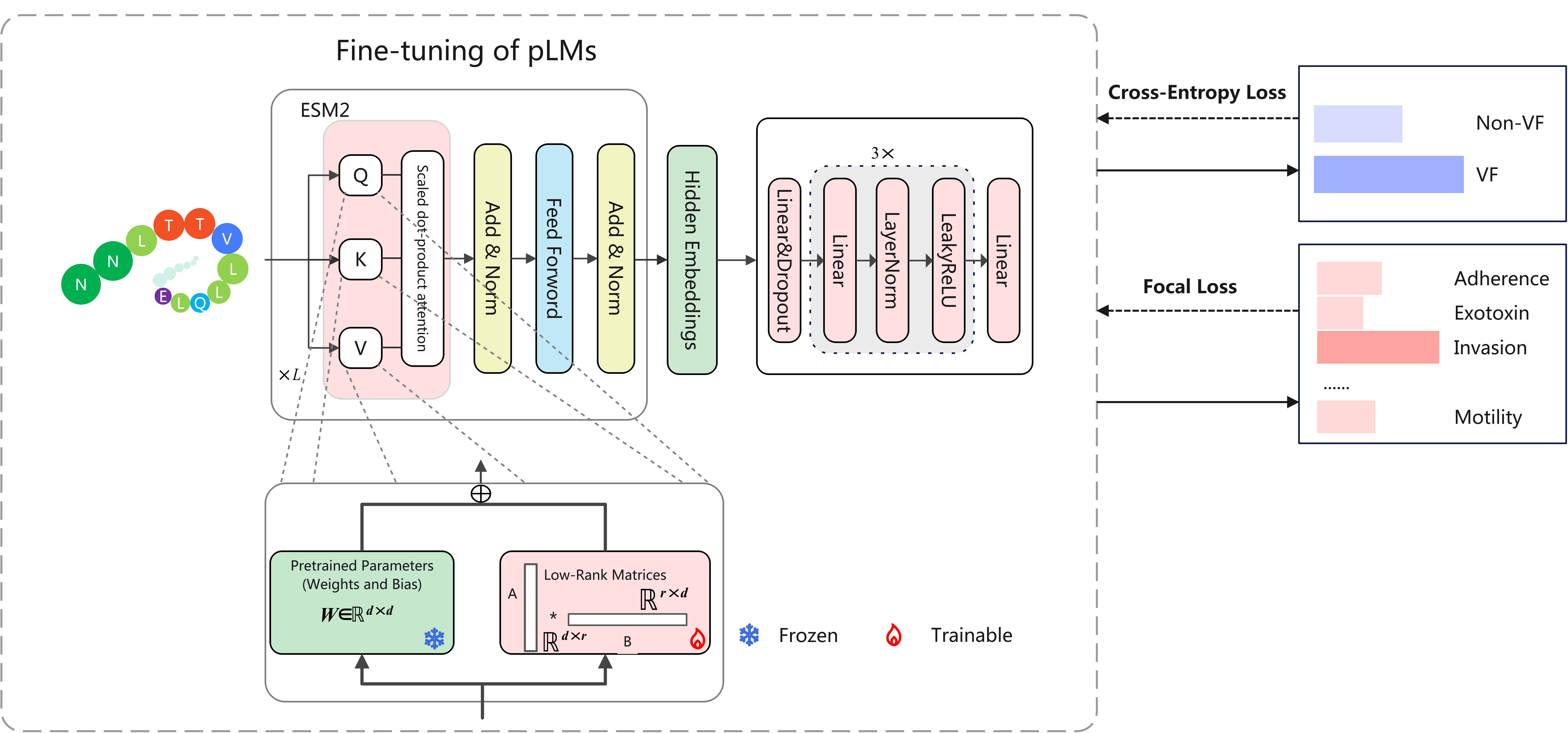
We present VirulentHunter, a novel deep learning framework for simultaneous VF identification and classification directly from protein sequences. We constructed a comprehensive, curated VF database by integrating diverse public resources and rigorously expanding VF category annotations. Benchmarking demonstrates that VirulentHunter significantly outperforms existing methods, particularly for VFs lacking detectable homology.
- python 3.8.13
- pytorch 2.4.1
- transformers 4.44.2
- biopython 1.83
- peft 0.7.1
To use the VirulentHunter codes, you first need to download the 'esm2_t30_150M_UR50D' model and put it under the fold of 'models/', and run the following command:
predict.py -i data/test.fasta -o results/predict_results.txt- Binary task:
python main.py --esm_path models/esm2_t30_150M_UR50D --input_fasta_path data/binary/ --input_label_path data/binary/ --max_len 2000- VF category task:
python main_multi_label.py --esm_path models/esm2_t30_150M_UR50D --input_fasta_path data/multi-label/train.fasta --input_label_path data/multi-label/train_labels.pkl --max_len 2000We are pleased to announce the launch of our web service, VirulentHunter, designed for the identification and prediction of virulence factors (VFs). The platform supports the input of protein sequences, strain genomes, and metagenomic data. VirulentHunter analyzes the input data and provides detailed outputs, including the identified virulence factors (if the query is classified as a VF) and their corresponding prediction probabilities. Visit the website at http://www.unimd.org/VirulentHunter to explore its capabilities. For users with large-scale data prediction needs, we recommend deploying a local version of the service to ensure efficient and customized processing.
[1] B. Liu, D. Zheng, S. Zhou, L. Chen, and J. Yang, “VFDB 2022: a general classification scheme for bacterial virulence factors,” Nucleic Acids Research, vol. 50, no. D1, p. D912, Jan. 2022, doi: 10.1093/nar/gkab1107.
[2] S. Sayers et al., “Victors: a web-based knowledge base of virulence factors in human and animal pathogens,” Nucleic Acids Research, vol. 47, no. D1, pp. D693–D700, Jan. 2019, doi: 10.1093/nar/gky999.
[3] . D. Olson et al., “Introducing the Bacterial and Viral Bioinformatics Resource Center (BV-BRC): a resource combining PATRIC, IRD and ViPR,” Nucleic Acids Research, vol. 51, no. D1, pp. D678–D689, Jan. 2023, doi: 10.1093/nar/gkac1003.
[4] L. Fu, B. Niu, Z. Zhu, S. Wu, and W. Li, “CD-HIT: accelerated for clustering the next-generation sequencing data,” Bioinformatics, vol. 28, no. 23, pp. 3150–3152, Dec. 2012, doi: 10.1093/bioinformatics/bts565.
[5] T. Hamamsy et al., “Protein remote homology detection and structural alignment using deep learning,” Nat Biotechnol, pp. 1–11, Sep. 2023, doi: 10.1038/s41587-023-01917-2.